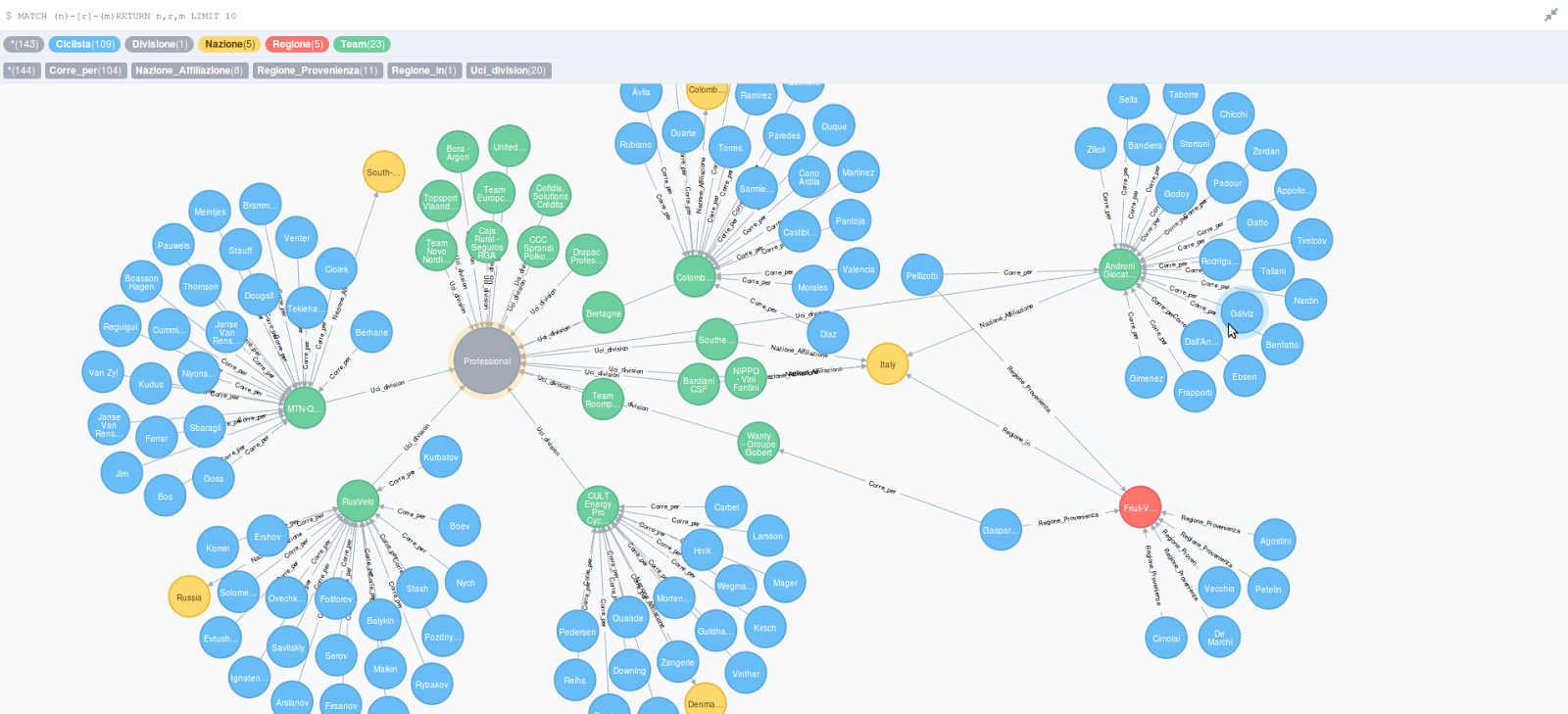
Hence, after processing all data, each GO term node holds the total number of times it has been assigned to a particular organism. Based on the InterProScan output, the number of times a GO term is found in a certain species is added as a field to the node representing that particular GO term. It is important to note that the GO database does not follow a tree structure but rather a more general Directed Acyclic Graph (DAG), i.e. 
The GO-basic data set was downloaded from the Gene Ontology Consortium website ( ), processed with a Python script and stored in a Neo4j graph database, with each node corresponding to a GO term and “ISA” relations as edges. 
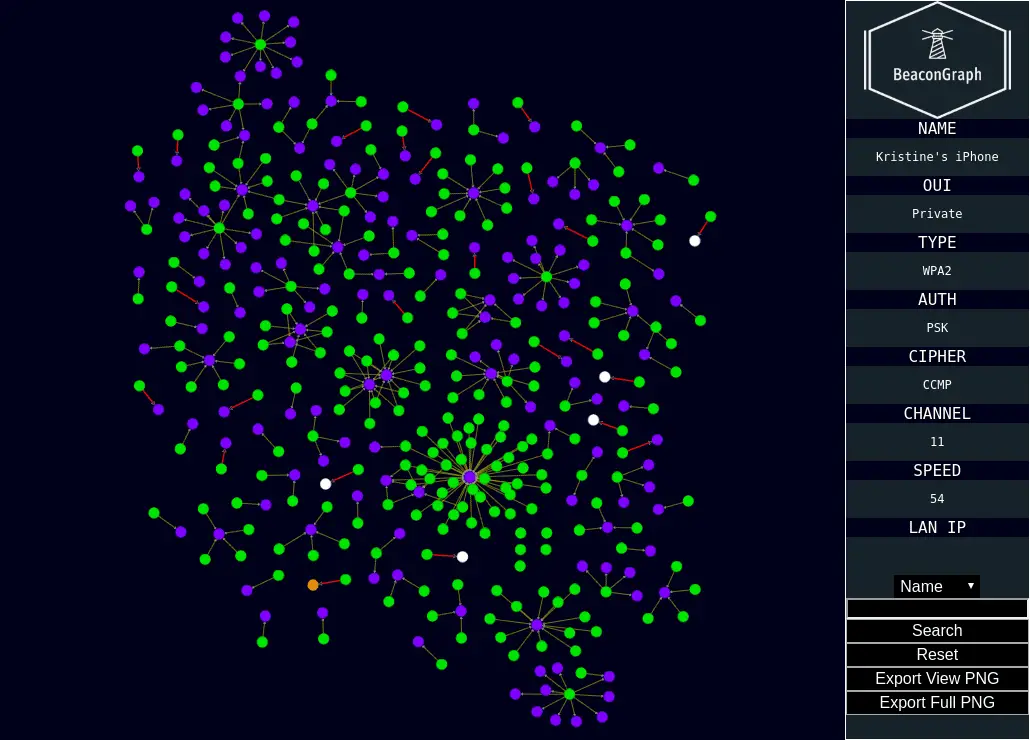
Here we outline the methodology underlying the approach. The applicability of this approach is demonstrated, by analyzing seventeen fungal species, including two Synchytrium endobioticum strains and six reference species, divided into four functional groups. After addition of InterProScan results, these graphs can be relatively easily queried to perform complex analyses on the data. Here, we present a new approach building on graph database technologies and Cytoscape to mine functional annotations, enable comparison of GO terms and KEGG pathways and visualize the results.īoth GO and KEGG databases store highly connected information, suitable for storing in a Neo4j graph database. However, there are currently no tools available to store, visualize and analyze the functional annotation of several species combined. These approaches mainly focus on comparing a single species of interest to one known reference species. Genes can be functionally annotated using InterProScan or Blast2GO, studying presence/absence of specific Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway elements or Gene Ontology (GO) enrichment. Combined with sequence information, these data can then be used for comparative genomics efforts to interpret the functional makeup of the organism under study to that of a related organism. It's like a skill tree for learningĪlso if you don't need to embed it in a webpage you can just use cytoscape.A first step following in silico reconstruction of a genome is usually to produce a structural annotation of the genome to identify repeats, genes and other genomic features. I built an interactive map for self-teaching online. Have you had a look at Cytoscape? I think this could work for what you're doing. The formatting stuff was all in Cytoscape, which is a pretty sick program for graph stuff.īest software for making figures and diagrams? Has anybody tried Cytoscape? () (DNS is SERVFAILing at the moment.) I use it for a combination of "no K" clustering (general exploration) and what's referred to in threat intelligence by the term of art "pivoting". I've been thinking that Gephi is getting long in the tooth.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
